


 |  |  |
As explored in Section 4.6, curves (Definition 45) are a versatile entity in the context of differential geometry as well as in physical applications as seen in Section 5.3. Geometric structures such as curves are defined on top of topological spaces, whose discrete representation has already been outlined in Section 6.3. The structure of fiber bundles as described in Section 6.4 allows for a simple storage of vector fields (Definition 62), which are intrinsically connected to their integral curves (Definition 63). This connected notion should be carried over from the continuous setting to the world of digital computers.
However, since the data items, such as vector fields, are stored only on discrete points, associated with the CW complex (Definition 34), a curve can also only be approximated. Returning to a very basic notion of the curve, as a mapping (Definition 20) associating a single parameter to a set of points in a manifold (Definition 37), it becomes apparent that an approximation of a curve can be achieved by storing a set of points with an additional mapping relating the CW complex and the curve.
Thus an integral curve, as a one-dimensional differentiable manifold, has its own representation, which is initially completely independent of the topological space it traverses. However, a mapping must exist and also be explicitly available, in order to use a curve within the discrete digital world. This mapping is readily found, when turning to atlases (Definition 36) and their charts. Thus by calculating the coordinates of points, it is possible to relate the points of the curve with its embedding manifold. However, in a manifold the obtained coordinates are valid only locally and worthless, since they degenerate to a mere collection of numbers, without knowledge of the chart in which they are obtained. A simple example is obtained by specifying points using barycentric coordinates within each of the cells. Without knowledge which cell to reference it is not possible to correctly locate the point in the CW complex. In the case of an affine space (Definition 79), the obtained coordinates have global scope and no further information needs to be specified, beside the knowledge that it is indeed an affine space.
The points comprising the curve can either be represented using a CW complex but can also be given in an implicit manner, using a prescription to determine points from a parameter. In the case of integral curves, the derivative with respect to the parameter of the mapping used for point calculations must match the given vector field. The simple yet powerful structure of polynomials (Definition 19) makes them well suited for this task. In order to avoid issues with high polynomial orders due to Runge’s phenomenon [114] interpolation is usually restricted to locally using low orders of polynomials, thus modelling the curve piecewise by several polynomials. This piecewise representation of a curve has become known as a spline.
Since the local nature of splines corresponds to the local nature of charts, they are well suited to directly provide coordinates for points of the curve, making them directly applicable in the previously described scheme.
The identification of points using coordinates seems to alleviate the problem of dealing with points belonging to two manifold structures, the solution suffers from imperfections due to the numerical issues connected with the calculation of the coordinates. It thus degrades the reliability of determining the identity of points, as it moves it from a problem related to set theory and topology to a geometric setting.
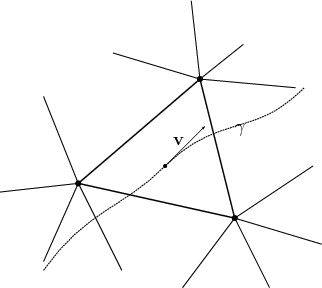
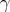 passes through several elements, thus as the curve’s parameter is varied
the location corresponds to different cells.
passes through several elements, thus as the curve’s parameter is varied
the location corresponds to different cells.In case the description uses local expressions, corresponding to cells, it is important to identify, when a curve moves from one cell to the next (illustrated in Figure 6.2), as it indicates the limit of its validity. Thus, the intersection of a curve with the boundary of a cell is worth to be investigated further.
In the case of a Riemannian base manifold and hence the availability of a mapping between 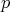 -forms
and vectors due to an isomorphism as indicated in Equation 4.156, the boundaries of tetrahedral and
hypercube cells can be geometrically represented using a normal vector
-forms
and vectors due to an isomorphism as indicated in Equation 4.156, the boundaries of tetrahedral and
hypercube cells can be geometrically represented using a normal vector  to the boundaries surface
using a simple formula
to the boundaries surface
using a simple formula

 represents a point to be tested.
represents a point to be tested.
Given the curve has a local polynomial representation in a parameter 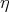

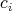 are connected to the current cell, e.g., derived from quantities
associated to this cell. Inserting Equation 6.4 into Equation 6.3, the intersections are obtainable
as
are connected to the current cell, e.g., derived from quantities
associated to this cell. Inserting Equation 6.4 into Equation 6.3, the intersections are obtainable
as


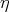 , whose coefficients are given by the inner product
of the curve parameters and the normal vector, whose roots indicate the possible intersections. The
correct intersection can then be chosen from the calculated roots.
, whose coefficients are given by the inner product
of the curve parameters and the normal vector, whose roots indicate the possible intersections. The
correct intersection can then be chosen from the calculated roots.
As a machine does not possess any intuitive notion which boundary component may be intersected and which not, the intersection needs to be evaluated for every boundary component (sub-cell) of the currently investigated cell. The boundary sub-cells and their face normals thereby need to be available from the base topological framework.
The following piece of code illustrates an implementation of the outlined algorithm using the GSSE infrastructure.
These data types are defined and evaluated at compile time, where DIM represents the global manifold
dimension and DIMB represents the corresponding facet sub dimension. The type CellTopology
describes the discrete cell representation of the underlying manifold as explicit mesh, e.g., a simplex
mesh or a cube grid, as outlined is Section 6.3. Finally, the ci need to be available or calculated form
values defined on the base topology, as sketched in Section 6.4, to represent the curve 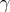 according to
Equation 6.4. Here a cubic polynomial, thus of degree
according to
Equation 6.4. Here a cubic polynomial, thus of degree 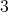 , is chosen for illustration. An application of
this procedure, where the coefficients
, is chosen for illustration. An application of
this procedure, where the coefficients 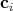 are determined from physical considerations, is available in
Section 7.2.1.
are determined from physical considerations, is available in
Section 7.2.1.
Where the previous snippet of code presented a compile time structure, the next piece of code is evaluated at run time. The new_cell along with the roots of the curve parameter are calculated from the given current cell, its boundaries and their geometry, and the ci as indicated by the global mesh structure.
The local description due to the use of splines limits the order of the used polynomials, thereby also limiting complications connected to the calculation of the roots of the polynomial. Once the next cell has been determined, the next polynomial of the spline corresponding to the new cell can be used.
 |  |  |